mRNA display is often described as one of the most powerful technologies for discovering high-diversity peptide binders. In principle, it offers extraordinary library sizes, tight genotype–phenotype linkage, and direct access to functional peptides. In practice, however, many labs never make it past the point of curiosity. Historically, that hesitation has not come from a single barrier, but from a combination of technical complexity and reliance on specialized techniques that are not universally accessible.
Radioactivity is often the most daunting of those barriers. Many of the foundational mRNA display experiments, including the seminal work by Roberts and Szostak1, relied on radiolabeled amino acids to monitor translation efficiency and peptide–mRNA fusion formation. These approaches are powerful, but they also require dedicated infrastructure, regulatory approvals, safety training, and operational experience that many modern discovery labs simply do not have. Reviews of the field, such as those by Kamalinia and colleagues2, highlight both the extraordinary power of mRNA display and the substantial experimental care and optimization required to implement it successfully. Even beyond detection of radiolabeled amino acids, the overall workflow can feel daunting to groups without prior hands-on experience.
When we set out to build an mRNA display platform at Parabilis, we were very much in that position ourselves. We were convinced of the value of the technology and excited by its potential to expand our Helicon discovery platform, but we did not have access to radioactive workflows, nor did we want that to become a gating factor. Rather than treating radioactivity as a prerequisite, we asked a more pragmatic question: how could we make peptide–mRNA fusions visible, quantifiable, and reliable using tools that are already accessible to most labs?
Our first instinct was to lean on techniques we were already familiar with. Immunoblotting and quantitative PCR (qPCR) are workhorses in molecular biology, and there is prior literature showing that both can be applied to mRNA display systems. Several groups have validated peptide–mRNA fusions using epitope tags such as FLAG3,6 and T75, while others have used qPCR to quantify fusion recovery or enrichment across selection rounds.4,7 That literature suggested a non-radioactive path was viable, but translating those demonstrations into a routine workflow still required innovation and iteration.
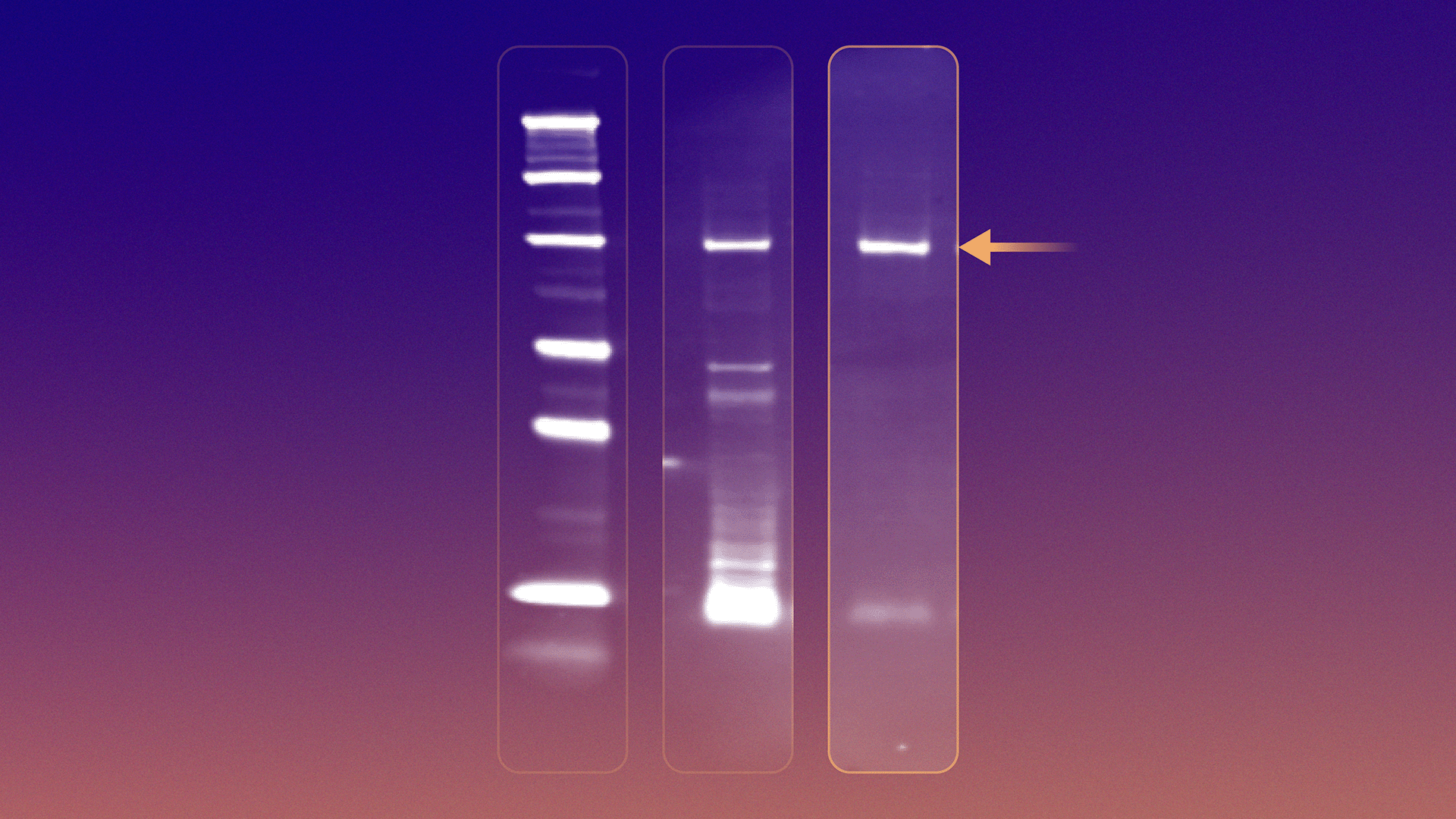
Figure 1 shows where we ultimately landed for qualitative detection. We explored several epitope tags to visualize peptide–mRNA fusions by immunoblotting. Some approaches worked only partially.
- His epitope tag detection, for example, proved unusable due to the abundance of His-tagged in vitro translation components, which overwhelmed the signal, and overall weak antibody engagement (Figure 1A).
- FLAG epitope tag detection was possible but, in our hands, the signal was weak and inconsistent, likely reflecting a combination of low fusion abundance and antibody performance.
- Consistent with a review on epitope tags8, the His epitope proved more useful as an affinity purification tag and the FLAG epitope proved insufficiently sensitive at a single copy.
An HA epitope tag ultimately emerged as the most reliable and sensitive solution (Figure 1B). With the right placement, HA-tagged peptide–mRNA fusions produced a clear, specific band at the expected molecular weight, both in crude translation mixtures and after purification. This gave us a fast, intuitive way to answer basic but essential questions: did translation work, did fusion occur, and did purification improve signal quality?
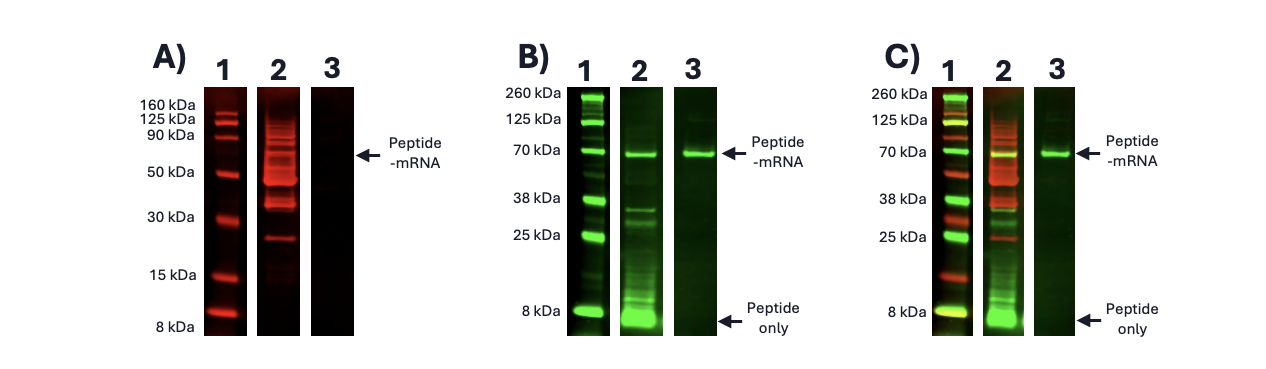
Figure 1. Visualization of mRNA displayed stapled peptide library. A) Anti-his immunoblot of tagged mRNA displayed stapled peptide library (70 kDa); B) Anti-HA immunoblot of tagged mRNA displayed stapled peptide library; C) Overlayed anti-his & anti-HA immunoblots of mRNA display stapled peptide library. 1 – Protein Ladder; 2 – Crude Translation; 3 – Purified peptide-mRNA library.
While immunoblotting solved the problem of visibility, it quickly became clear that it was not the right tool for quantification. Estimating library size or fusion yield from band intensity alone was inconsistent and highly sensitive to experimental conditions (Figure 2A). For that, we turned to qPCR. By reverse-transcribing the mRNA component of the fusion and quantifying the resulting cDNA, we could indirectly but sensitively measure the number of peptide–mRNA molecules present. As shown in Figure 2B, this approach produces clean standard curves with excellent linearity and enables routine estimation of library sizes in the 10¹³–10¹⁴ molecule range. Importantly, this approach can be done without requiring specialized infrastructure.
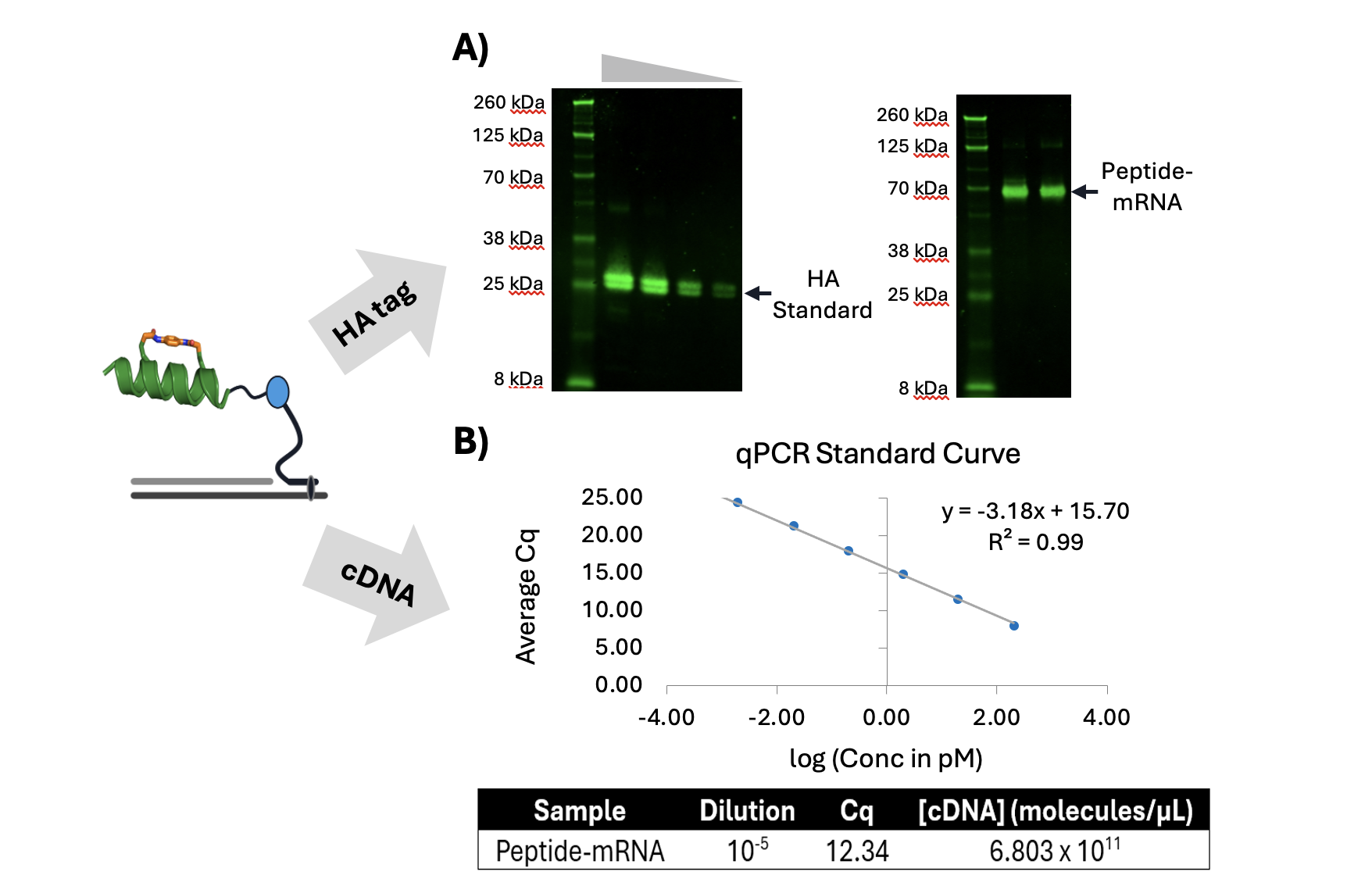
Figure 2. Visualization and quantification of mRNA displayed stapled peptide. A) Anti-HA immunoblot of HA-tagged mRNA displayed stapled peptide (70 kDa) and HA-tag standard (21 kDa). HA-tag standard previously shown to run closer to 25 kDa on 4-12% Bis-Tris polyacrylamide gel.; B) Quantitative PCR (qPCR) standard curve and quantification of cDNA containing mRNA displayed stapled peptide library.
Together, immunoblotting and qPCR gave us a complementary, radioactivity-free detection framework. Immunoblotting provides a qualitative, confidence-building readout of fusion formation and integrity. qPCR provides a quantitative handle on library size and recovery. Combined, these methods meet the core needs of an mRNA display workflow in a way that is accessible, scalable, and reproducible.
Looking back, the most important lesson for us was not tied to any single technique. It was the realization that building an mRNA display platform does not require reproducing every aspect of legacy workflows. By acknowledging our constraints up front and leaning into tools we already understood, we were able to adapt a powerful technology to our environment rather than reshaping our lab around it. For teams curious about mRNA display but hesitant due to perceived barriers, we hope this story makes the path feel more navigable. You do not need radioactive amino acids to get started. You need clear questions, familiar tools, and a willingness to innovate and iterate.
Authored by Justin Lawrie, Platform Technology team member
References
1. Roberts RW, Szostak JW. RNA-peptide fusions for the in vitro selection of peptides and proteins. Proc Natl Acad Sci U S A. 1997 Nov 11;94(23):12297-302. doi: 10.1073/pnas.94.23.12297. PMID: 9356443; PMCID: PMC24913.
2. Kamalinia G, Grindel BJ, Takahashi TT, Millward SW, Roberts RW. Directing evolution of novel ligands by mRNA display. Chem Soc Rev. 2021 Aug 21;50(16):9055-9103. doi: 10.1039/d1cs00160d. Epub 2021 Jun 24. PMID: 34165126; PMCID: PMC8725378.
3. Ueno S, Arai H, Suzuki M, Husimi Y. An mRNA-protein fusion at N-terminus for evolutionary protein engineering. Int J Biol Sci. 2007 Aug 29;3(6):365-74. doi: 10.7150/ijbs.3.365. PMID: 17848981; PMCID: PMC1975775.
4. Villequey C, Zurmühl SS, Cramer CN, Bhusan B, Andersen B, Ren Q, Liu H, Qu X, Yang Y, Pan J, Chen Q, Münzel M. An efficient mRNA display protocol yields potent bicyclic peptide inhibitors for FGFR3c: outperforming linear and monocyclic formats in affinity and stability. Chem Sci. 2024 Mar 18;15(16):6122-6129. doi: 10.1039/d3sc04763f. PMID: 38665530; PMCID: PMC11040643.
5. Muranaka N, Hohsaka T, Sisido M. Four-base codon mediated mRNA display to construct peptide libraries that contain multiple nonnatural amino acids. Nucleic Acids Res. 2006 Jan 5;34(1):e7. doi: 10.1093/nar/gnj003. PMID: 16397292; PMCID: PMC1325208.
6. Doshi R, Chen BR, Vibat CR, Huang N, Lee CW, Chang G. In vitro nanobody discovery for integral membrane protein targets. Sci Rep. 2014 Oct 24;4:6760. doi: 10.1038/srep06760. PMID: 25342225; PMCID: PMC4208029.
7. Otero-Ramirez ME, Matoba K, Mihara E, Passioura T, Takagi J, Suga H. Macrocyclic peptides that inhibit Wnt signalling viainteraction with Wnt3a. RSC Chem Biol. 2020 Mar 24;1(1):26-34. doi: 10.1039/d0cb00016g. PMID: 34458746; PMCID: PMC8382136.
8. Terpe K. Overview of tag protein fusions: from molecular and biochemical fundamentals to commercial systems. Appl Microbiol Biotechnol. 2003 Jan;60(5):523-33. doi: 10.1007/s00253-002-1158-6. Epub 2002 Nov 7. PMID: 12536251.